Phosphorylation of unique domains of Src family kinases
Members of the Src family of kinases (SFKs) are non-receptor tyrosine kinases involved in numerous signal transduction pathways. The catalytic, SH3 and SH2 domains are attached to the membrane-anchoring SH4 domain through the intrinsically disordered “Unique” domains, which exhibit strong sequence divergence among SFK members.
In the last decade, structural and biochemical studies have begun to uncover the crucial role of the Unique domain in the regulation of SFK activity. This mini-review discusses what is known about the phosphorylation events taking place on the SFK Unique domains, and their biological relevance.
The modulation by phosphorylation of biologically relevant inter- and intra- molecular interactions of Src, as well as the existence of complex phosphorylation/dephosphorylation patterns observed for the Unique domain of Src, reinforces the important functional role of the Unique domain in the regulation mechanisms of the Src kinases and, in a wider context, of intrinsically disordered regions in cellular processes.
The Src kinase family is composed of 10 proteins: Src, Frk, Lck, Lyn, Blk, Hck, Fyn, Yrk, Fgr, and Yes. Src family kinases (SFKs) are membrane-associated, non-receptor tyrosine kinases that act as important signaling intermediaries regulating a variety of outputs, such as cell proliferation, differentiation, apoptosis, migration, and metabolism.
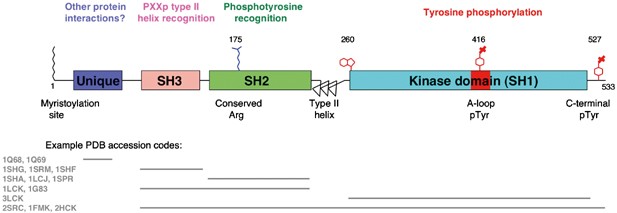
All SFKs share the same domain arrangement: a large catalytic C-terminal domain is preceded by regulatory Src 2 and 3 homology domains (SH2 and SH3, respectively) and the membrane-anchoring SH4 N-terminal region, which contain myristoylation and palmitoylation sites as well as positively charged residues.
The SH3 and SH4 domains are linked by an intrinsically disordered segment of 50–90 residues, called the Unique domain (UD) because of the lack of sequence similarity among the different SFKs. However, the UD of each individual SFK member is well conserved between different organisms suggesting a more specific role than that of a simple spacer.
